|
12/8/2023 0 Comments Youtube r studio anova
This script uses an extension of vegan library's bioenv() function and finds the best set of environmental variables with maximum (rank) correlation with community dissimilarities and then plots them as vectors along with the best subset of taxas on the NMDS plot. The above code will produce the following plot: P <-p +geom_point (aes ( shape= Depth ) ) +scale_shape_manual (values=shape_values ) +theme_bw ( ) pdf ( "NMDS.pdf" ) print (p ) dev.off ( ) P <-p + geom_path ( data=df_ell, aes (x=NMDS1, y=NMDS2 ), size= 1, linetype= 2 ) P <-p + annotate ( "text" ,x=an $x ,y=an $y ,label=an $group ,size= 4 ) 
P <- ggplot ( data=NMDS ,aes (x ,y ,colour=Country ) ) # = # Tutorial on drawing an NMDS plot using ggplot2 # by Umer Zeeshan Ijaz () # =Ībund_table head(grouping_info) # X1 X2 X3 # T_2_1 T 2 1 # T_2_10 T 2 10 # T_2_12 T 2 12 # T_2_2 T 2 2 # T_2_3 T 2 3 # T_2_6 T 2 6 #Load vegan library library ( vegan ) #Get MDS stats This script finds a non-parameteric monotonic relationship between the dissimilarities in the samples matrix, and plots the location of each site in a low-dimensional space (similar to principle component analysis). Please make sure that you have the necessary packages installed! You can then click on "Session" on the menu to set working directory to the source file location, and click on the "source" button to run the analysis as shown below: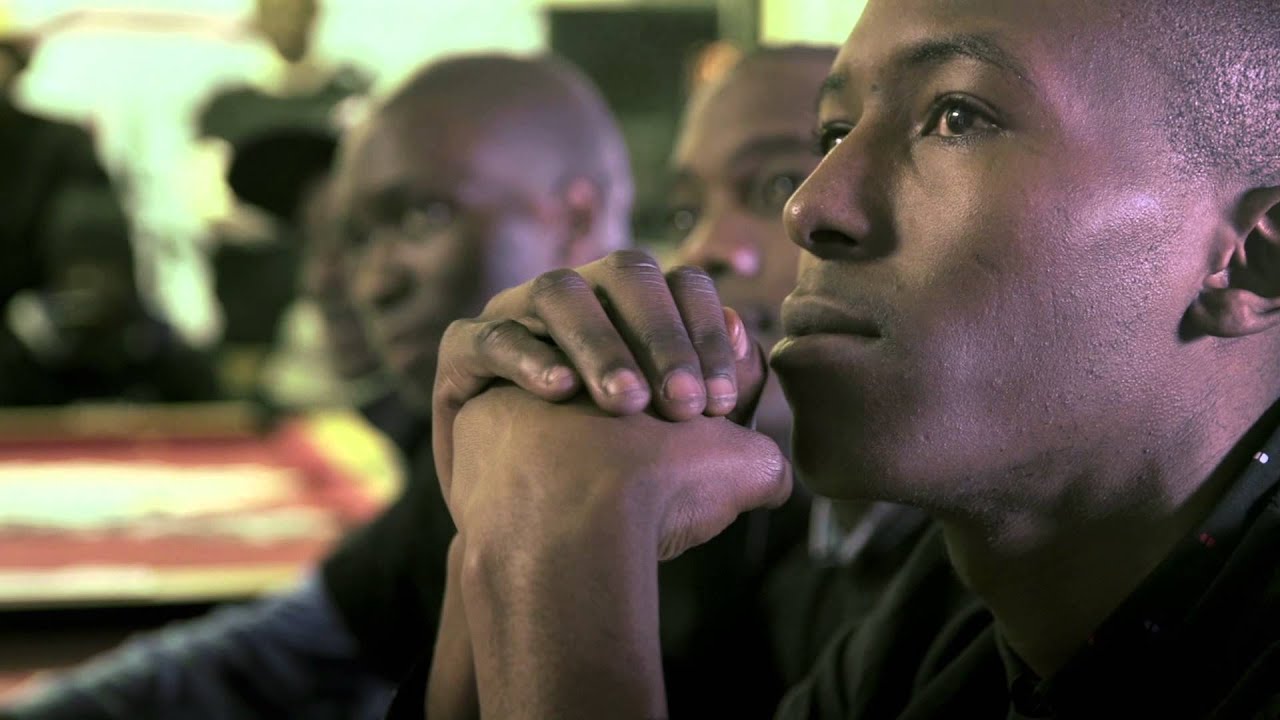 
SPE_pitlatrine.csv (species abundance file)Īll_Good_P2_C03_Taxonomy.csv (OTUs taxonomic breakdown)Īll_Good_P2_C03.tre (OTUs phylogenetic tree in newick format) Instructionsĭownload RStudio, copy the files listed above in a folder, load the IDE, paste the scripts as they are on the left-pane and save them in the same folder. Please note that in this format, any categorical information is reflected in the sample names, and the metadata contains the continuous environmental measures. If you format your tables in the same manner as I have, you will need very little editing of the scripts to perform analyses of your datasets. In the files given below, I have used the following naming convention for sample names. 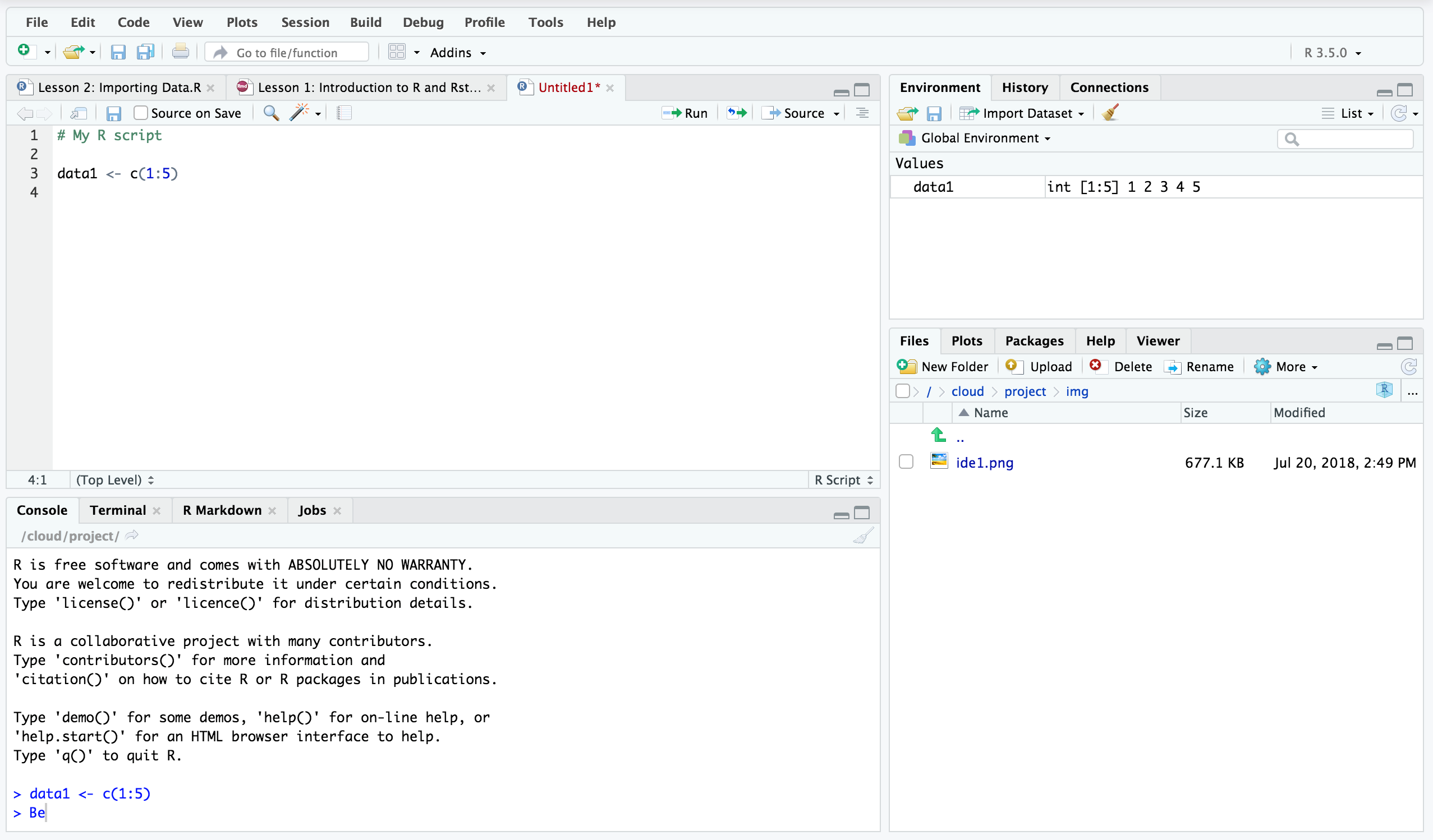
To test my code, you can use the toy pitlatrine dataset which is generated by metagenomic 16SrRNA sequencing of latrines from Tanzania and Vietnam at different depths (multiples of 20cm). Please cite the following paper if you find the code useful:ī Torondel, JHJ Ensink, O Gundogdu, UZ Ijaz, J Parkhill, F Abdelahi, V-A Nguyen, S Sudgen, W Gibson, AW Walker, and C Quince.Īssessment of the influence of intrinsic environmental and geographical factors on the bacterial ecology of pit latrines R code for ecological data analysis R code for ecological data analysisīy Umer Zeeshan Ijaz Material ggplot2.pdf
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
